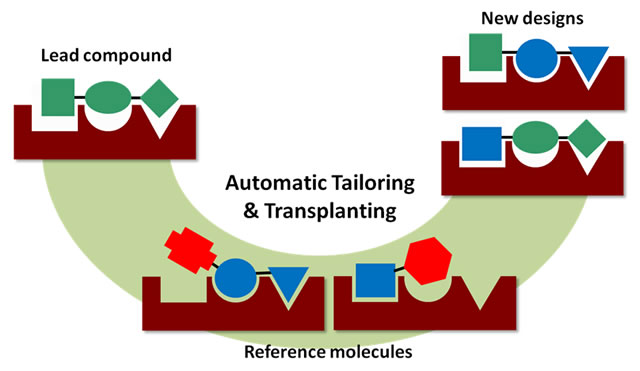
The Automatic Tailoring and Transplanting (AutoT&T) method is developed as a versatile computational tool for automatic lead optimization as well as lead discovery in molecular-targeted drug design. This method detects suitable fragments on reference molecules and then transplants them onto the given lead compound to generate new ligand molecules. Binding affinities, synthetic feasibilities, and drug-likeness properties are also evaluated to select the promising candidates for further consideration.
- AutoT&T does not reply on a pre-defined built-in library of building blocks. All fragments used in structural operation are truncated directly from reference molecules. Reference molecules are supplied by the users in a straightforward manner.
- AutoT&T does not need to perform conformational sampling during structural construction. The binding poses of all reference molecules are generated in prior, for example, through a virtual screening job against the given target protein.
- AutoT&T is able to conduct a systematic crossover between the lead molecule and all given reference molecules. It does not need to tackle the “combinatorial explosion” problem confronted by conventional build-up methods. We have tested AutoT&T on several different types of drug targets. Our results indicated that AutoT&T could not only produce more reasonable designs, it was also much more efficient than the conventional build-up methods.
AutoT&T version 2
The original version of AutoT&T was published in 2011. Significant improvements have been made since then. The new version, i.e. AutoT&T2, is faster by up to a few thousand folds in a multi-round optimization job. New applications are also implemented in AutoT&T2, for example, performing structural crossover among multiple lead molecules or design of ligand molecules from scratch. Please refer to our latest publication or the user manual for more details.
Click this link for a copy of the user manual of AutoT&T2, [User manual of AutoT&T2]
The AutoT&T2 software suite is now available under a modest license fee. The license fee is collected for non-profit purpose, e.g. maintenance and personnel cost. Different license fees apply to different categories of users:
- (1) The license fee for all academic, governmental, and other non-profit organizations is 300 USD (2000 CNY for users in China).
- (2) The license fee for all commercial, industrial, and other for-profit organizations is 3000 USD (20000 CNY for users in China). This fee grants a site license with an unlimited number of users.
- (3) The license fee for non-profit and for-profit computing centers is also 3000 USD (20000 CNY for users in China).
To obtain the AutoT&T2 software suite, please go to the following [REGISTRATION] page, select the correct category (i.e. academic/government or commercial) and fill in necessary contact information. Upon registration, an end-user license agreement must be signed for copyright protection. Then, we will provide you a formal invoice with payment information. Once we receive the payment, we will send you through e-mail further instruction on downloading the AutoT&T2 software suite.
The AutoT&T2 suite includes the source codes and executable codes (for Linux and Windows platforms) of all major functions, the user manual, and the necessary material for running the several demo examples described in our publication. It is provided as a TAR.gz package (155 MB in size).
Reference:
- Yan Li, Zhixiong Zhao, Zhihai Liu, and Renxiao Wang*, "AutoT&T v2: An Efficient and Versatile Tool for Lead Structure Generation and Optimization", J. Chem. Inf. Model., 2016, 56, 435-453. (AutoT&T version 2)
- Yan Li, Yuan Zhao, Zhihai Liu, Renxiao Wang*, "Automatic Tailoring and Transplanting: A Practical Method that Makes Virtual Screening More Useful", J. Chem. Inf. Model.,2011, Vol. 51, pp 1474-1491. (AutoT&T version 1)